Oncology
Abstract
BACKGROUND. Asparaginase is essential for curing acute lymphoblastic leukemia (ALL), but its use is limited by asparaginase-associated pancreatitis (AAP), a severe and unpredictable toxicity lacking validated prospective biomarkers. We sought to define early systemic molecular features of susceptibility to AAP. METHODS. We performed longitudinal lipidomic and proteomic profiling in two independent pediatric ALL cohorts (n = 161; 79 AAP cases, 82 controls) using paired blood samples collected before asparaginase exposure and at the end of induction therapy (including a single dose of asparaginase), thereby capturing pre-injury biology rather than consequences of pancreatitis. We applied differential abundance and network-based analyses, and integrated lipid–cytokine associations using proteomics. RESULTS. Across cohorts, we identified a reproducible lysophosphatidylcholine (LPC)–centered signature characterized by attenuated induction therapy-associated LPC responses and disruption of LPC co-regulation at the network level. Proteomic profiling revealed enrichment of cytokine signaling pathways, and integrative analyses demonstrated altered lipid–cytokine coupling, including a flip in association direction for LPC species and interleukin-18 (IL-18) between cases and controls. Although IL-18/LPC ratios do not differ globally, elevated post-induction IL-18/LPC ratios identify AAP risk within a protocol-defined very high-risk ALL subgroup (AUC = 0.81). CONCLUSION. These findings support a systems-level model in which failure of coordinated lipid–immune responses under therapeutic stress confers vulnerability to AAP, providing a framework for validation and mitigation strategies. TRIAL REGISTRATION. NCT00400946; NCT01574274; NCT03020030 (parent trials). FUNDING. Servier Pharmaceuticals (IIT-95014-027-USA); SDRC (P30DK116074); Stanford SPARK; Fonds de Recherche du Québec – Santé; Fondation Charles-Bruneau; The Leukemia & Lymphoma Society of Canada.
Authors
Cheng-Yu Tsai, Na Bo, Thai Hoa Tran, Maisam Abu-El-Haija, Gayathri Swaminathan, Bomi Lee, Sudhir Ghandikota, Li Wen, Yves Théorêt, Steven D. Mittelman, Elena J. Ladas, Anil G. Jegga, Lewis B. Silverman, Ying Ding, Sohail Z. Husain
Abstract
Immune checkpoint inhibitors have transformed cancer therapy, yet many patients fail to achieve durable responses due to insufficient T cell reinvigoration. Cytokines offer promise for enhancing immunotherapy, but their clinical use is limited by toxicity and a narrow therapeutic index. Immunocytokines, engineered fusion proteins combining antibody specificity with cytokine activity, aim to overcome these challenges by targeting cytokine delivery to immune cells or the tumor microenvironment. We describe SAR445877 (SAR’877), a novel PD-1-targeted immunocytokine that fuses a high-affinity anti-PD-1 antibody with a detuned IL-15/IL-15Rα sushi domain complex. SAR’877 blocks PD-1/PD-L1 and PD-1/PD-L2 interactions while selectively delivering IL-15 signals to PD-1+ T cells, enhancing proliferation and activation of antigen-experienced CD8+ and CD4+ T cells and NK cells, while minimizing systemic inflammation. Mechanistically, SAR’877 activates STAT5 signaling in PD-1+ lymphocytes and restores effector function in exhausted T cells. In preclinical models, a murine surrogate of SAR’877 accelerated viral clearance and induced robust anti-tumor immunity by expanding cytotoxic CD8+ T cells and promoting Th1 polarization. Notably, SAR’877 outperformed anti-PD-1 plus untargeted IL-15, highlighting the therapeutic potential of targeted IL-15 delivery. These findings position SAR’877 as a promising next-generation immunotherapy with enhanced efficacy and reduced cytokine-associated toxicities.
Authors
Isaraphorn Pratumchai, Marie Bernardo, Julien Tessier, Jaroslav Zak, Kristi L. Marquardt, Joon Sang Lee, Maheeka Bimal, AHyun Choi, Anthony M. Byers, Mikielia G. Devonish, Roberto Carrio, Dan Lu, Stella A. Martomo, Jeegar Patel, Yu-an Zhang, Ingeborg M. Langohr, Virna Cortez-Retamozo, Dinesh S. Bangari, Angela Hadjipanayis, Xiangming Li, Valeria R. Fantin, Donald R. Shaffer, John R. Teijaro
Abstract
Tumor-infiltrating CD8 cells recognize neoantigens created by tumor-specific mutations. Nonetheless, even after checkpoint inhibitor therapy, most patients progress. A deeper understanding of anti-tumor responses could facilitate development of better therapies. To enable such studies, we applied TCXpress, a high throughput platform that clones fully expressible TCRs from single cells into retro- or lenti- viral vectors without sequencing or gene synthesis, to study TCRs from CD8 cells infiltrating mouse MC38 tumors. We expressed cloned TCRs in reporter cells and interrogated TCR specificity by coculturing them with B6WT3 cells transduced with tandem minigenes encoding predicted neoantigens. We isolated TCRs reactive against epitopes from mutant Rpl18, Adpgk, Psmd2, and Zc3h7b along with self-reactive TCRs that recognized normal B6 and MC38 cells. Importantly, we successfully treated MC38-bearing mice with T cells transduced with anti-Rpl18 TCRs. These results establish a system that could be used to study many types of T cell responses and validates a therapeutic approach that could be tested in the clinic.
Authors
Alexander M. Rowe, Smriti Chaurasia, Wenzhong Wei, Laura García-Diéguez, Katherine Querry, Johnathon G. Schiebel, Christy Smolak, Alexander G. Muralles, Daniel Wikenheiser, Kevin Quann, Collin Pirner, Kentin Codispot, Mark J. Shlomchik, Warren D. Shlomchik
Abstract
Germline BRCA1/2 pathogenic variant (PV) carriers have elevated young-onset breast cancer risk. To define the pretreatment genomic landscapes of young-onset gBRCA-associated breast cancer, we evaluated 136 treatment-naïve tumors diagnosed before age 50 (92.6% ≤40): gBRCA1 86(63.2%); gBRCA2 50(36.8%) in the prospective POSH study, and 66 noncarriers from The Cancer Genome Atlas. Using whole exome sequencing, we analyzed somatic variation, allele-specific loss of heterozygosity (asLOH), homologous recombination deficiency (HRD), and single-base substitution signatures (SBS). gBRCA1(93%) and gBRCA2(96%) breast cancers had high rates of asLOH, but differed significantly in average HRD scores (57.4 ± 1.3 vs 43.7 ± 1.5, P < 0.0001) and median SBS composition (%): SBS1 (aging-associated) 12.9 vs 7.3, P = 0.013; SBS18 (reactive oxygen species [ROS]-associated) 1.4 vs 0, P = 0.007; and SBS3 (HRD-associated) 27.3 vs 42.6, P = 0.002. Compared to gBRCA2 tumors, gBRCA1 tumors with asLOH were significantly enriched for alterations in Hallmark ROS, DNA repair, and epithelial-mesenchymal transition pathways. In ER-positive, HER2-negative tumors from gBRCA1/2 carriers compared to noncarriers, we found significant enrichment of RB1 (OR:6.3;95%CI:2.8–15.4;padj = 0.001), TP53 (OR:4.6;95%CI:1.9–12.1;padj = 0.017), FAT1 (OR:3.9;95%CI:1.84–8.7;padj = 0.013), and MYC (OR:4.0;95%CI:1.8–9.1;padj = 0.017) SNV/indels/CNVs, associated with CDK4/6i resistance. Together, these findings demonstrate significant differences between gBRCA1 and gBRCA2-associated breast cancers, and preexisting CDK4/6i resistance mechanisms supporting prospective trials with individualized therapy for gBRCA1 vs gBRCA2 carriers, and comparing PARPi to CDK4/6i for ER-positive gBRCA1/2-associated breast cancer.
Authors
Mwangala P. Akamandisa, Mingyi Xia, Wilson Cheah, Bradley Wubbenhorst, Kurt P. D'Andrea, Mengyao Fan, Jake S. Shilan, Dana Pueschl, Anupma Nayak, Hayley McKenzie, William Tapper, Ellen R. Copson, Ramsey I. Cutress, Susan M. Domchek, Diana M. Eccles, Katherine L. Nathanson
Abstract
Background Cancer accounts for over 20% of late post-transplant mortality, yet the contribution of genetic susceptibility to post-transplant cancer risk remains unclear. This study investigates germline genetic risk factors for post-transplant cancer in the Finnish population using data from the FinnGen cohort. Methods A pan-cancer polygenic risk score (PRS) was constructed using genetic variants identified in UK and US populations to assess the influence of common germline variants on time to first cancer diagnosis in 1,802 Finnish kidney transplant recipients (KTRs), of whom 317 developed post-transplant cancer. The PRS was first validated in the FinnGen non-transplantation cohort and subsequently applied to KTRs, with replication in lung and liver transplant recipients (n = 476). Functional relevance was explored by assessing associations between the PRS and expression levels of 2,923 plasma proteins in the UK Biobank (n = 53,013). Results Compared to a matched non-transplantation cohort (n = 68,294), KTRs exhibited earlier cancer onset. The PRS was significantly associated with time to first cancer diagnosis in the non-transplantation population (HR 1.04; 95% CI 1.038-1.056; p = 3.75 x 10-25). Among KTRs younger than 40 years, higher PRS was associated with earlier cancer onset (HR, 1.08; 95% CI ,1.01-1.17; p = 0.036), indicating a stronger genetic effect at younger ages. The PRS significantly (Bonferroni < 0.05) altered the regulation of 87 plasma proteins, several of which were known cancer-related markers. Conclusion Inherited genetic predisposition, captured by pan-cancer PRS, may contribute to individual susceptibility to cancer after solid organ transplantation, particularly at younger ages.
Authors
Jarmo Ritari, Kati Hyvärinen, Kirsi Jahnukainen, Jukka Partanen, Ilkka Helanterä, Timo Jahnukainen
Abstract
Merkel cell carcinoma (MCC) is a neuroendocrine carcinoma of the skin characterized by poor prognosis. This study aimed to explore the relationship between genetic alterations, tumor mutational burden (TMB), and MCC-specific survival (MCC-SS) in patients who underwent genomic profiling of tumors with OncoPanel. Univariate and multivariable analysis were used to assess the impact of genetic alterations on MCC-SS. Of the 188 identified patients, 164 were included in the analysis. The cohort had a mean age of 72.4 years (SD = 11.03), including 61.6% male. The median TMB was 5.32 (IQR = 3.04–25.53). Kaplan-Meier curves by high versus low TMB were significantly different (log-rank test, P = 0.017). PIK3CA (adjusted P = 0.003), SETBP1 (adjusted P = 0.002), KDR (adjusted P = 0.028), and RET (adjusted P = 0.033) were selected for multivariable analysis. In the multivariable regressions, only PIK3CA (HR = 2.07 [95% CI, 1.10–3.88]; P = 0.024) remained significant. PIK3CA remained significant across prespecified sensitivity analyses. In this study, high TMB and PIK3CA alterations were associated with poor MCC-SS. Identifying a higher-risk subgroup may inform risk stratification and motivate further evaluation of PI3K pathway targeting in future studies.
Authors
Matheus Lobo, Furkan Bahar, Julia L. Schnabel, Joao P. Duprat Neto, Aniket Shetty, Karam Khaddour, Manisha Thakuria, Ann W. Silk, James A. DeCaprio
Abstract
BACKGROUND. IL-7 is a critical cytokine in T cell development, survival, and homeostasis. Previous preclinical and clinical studies reported that IL-7 treatment increased T cell counts, but its effect on peripheral blood T cells in cancer patients and molecular mechanisms have not been explored. METHODS. We investigated effects of long-acting recombinant human interleukin-7 (rhIL-7-hyFc) on peripheral T cells in patients with advanced solid tumors. Peripheral blood samples were collected before and after treatment, followed by analysis through single-cell transcriptomics and flow cytometry. RESULTS. We found that rhIL-7-hyFc induced marked expansion of proliferating T cells, and promoted transcriptional changes associated with immune activation, cell cycle progression, and anti-apoptosis. Trajectory analysis revealed that post-treatment T cells had distinct transcriptional states enriched for cytokine- and TCR-mediated signaling pathways. Notably, a second dose administered after three weeks yielded diminished proliferation and minimal transcriptional changes, which were independent of antidrug antibody or CD127 downmodulation. Examination of elements of the IL-7 signaling pathway revealed intact proximal signaling (e.g., STAT5 phosphorylation) but downregulation of distal elements, including PIM-1 kinase and c-Myc. CONCLUSIONS. Our results demonstrate that rhIL-7-hyFc induces robust peripheral T-cell expansion and activation in patients with solid tumors, supporting its potential use for lymphopenic patients treated with cancer immunotherapy. TRIAL REGISTRATION. NCT03478995, NCT03619239. FUNDING. NRF-2022R1A2C3007292, RS-2024-00439160, RS-2025-02213409, RS-2025-25460003
Authors
Ho Cheol Jang, Jeong Yeon Kim, Sojeong Kim, Heewon Kim, Mi-Sun Byun, Myung Ah Lee, Jong Hee Chang, Do-Hyun Nam, Tae Won Kim, Sin-Soo Jeun, Joohyuk Sohn, Su-Hyung Park, Eui-Cheol Shin
Abstract
Inactivating NOTCH1 mutations in head and neck squamous cell carcinoma (HNSCC) were described over a decade ago, suggesting a tumorsuppressor function—unlike its oncogenic role in other tumors. Today, much debate persists regarding a putative oncogenic role in HNSCC as well, with reports that NOTCH1 signaling drives tumor growth and a cancerstemcell (CSC) phenotype. In this work, comprehensive experiments unequivocally demonstrate that NOTCH1 is a tumor suppressor in HNSCC regardless of mutation or activation status and that it reduces CSC frequency. We developed a signature of NOTCH1 activation showing the pathway is associated with very early differentiation, an altered tumor microenvironment, and better prognosis. Clarifying whether NOTCH1 occasionally functions as an oncogenic driver in HNSCC is crucial to prognosis and personalized therapy. The results presented unify the field, reconcile conflicting data, and provide critical insights into the biological and clinical significance of NOTCH1, with broader implications in other squamous carcinomas with NOTCH1 mutations.
Authors
Chenfei Huang, Shhyam Moorthy, Qiuli Li, Kazi M. Ahmed, Kalil Saab, Defeng Deng, Jiping Wang, Xiayu Rao, Jiexin Zhang, Yuanxin Xi, Jing Wang, Zhiyi Liu, Noriaki Tanaka, David A. Wheeler, Eve Shinbrot, Rami Saade, Curtis R. Pickering, Tong-Xin Xie, Adel K. El-Naggar, Abdullah A. Osman, Kunal Rai, Patrick A. Zweidler-McKay, John V. Heymach, Lauren A. Byers, Faye M. Johnson, Vlad C. Sandulache, Jeffrey N. Myers, Pedram Yadollahi, Mitchell J. Frederick
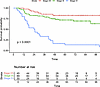
Loss of Tumor-Infiltrating Lymphocytes and Poor Response to Immunotherapy in IDH GOF Mutant Melanoma
Abstract
Recent innovations in melanoma treatment with immune checkpoint blockade (ICB) have improved overall outcomes for patients, however over 50% of patients still develop resistance to treatment. These patients either have intrinsic resistance, and never respond to therapy, or develop acquired resistance months or years into treatment. The mechanisms underlying ICB resistance remain poorly understood. Our data shows that isocitrate dehydrogenase gain of function (IDH GOF) mutant melanoma patients have a worse response to anti-PD1 immunotherapy. IDH mutations have been found to be oncogenic and associated with differential methylation in multiple cancers but are not yet characterized in human melanoma. Here, we investigate the clinical, immune, and transcriptional phenotypes of IDH GOF melanomas through analyses of clinical response, single-cell RNA sequencing, bulk RNA sequencing, and DNA methylation data. Single-cell data analysis shows decreased immune infiltrate and activity in the IDH GOF tumors. Bulk sequencing data demonstrates the association between IDH mutation, immune exclusion, and disruptions in global DNA methylation. The melanoma-derived genomic data presented supports previously described resistance mechanisms of IDH mutation in other cancer types and is the first demonstration of the role of IDH GOF in the human melanoma tumor microenvironment.
Authors
Emma Specht, Lakshmi Pakanati, Meng-Ju Wu, Russell W. Jenkins, Derek N. Effiom, Nabeel Bardeesy, Bradley E. Bernstein, Moshe Sade-Feldman, Christine G. Lian, Genevieve M. Boland, Elena Torlai Triglia, Sonia Cohen
Abstract
Supernumerary centrosomes are a hallmark of cancer. To maintain viability, cancer cells cluster these centrosomes during mitosis, enabling bipolar division similar to that of normal cells. Disruption of this centrosome clustering leads to multipolar anaphase and apoptosis (anaphase catastrophe), which selectively eliminates cancer cells harboring supernumerary centrosomes. In this context, because the motor protein KIFC1 contributes to centrosome clustering, we investigated whether targeting of this mechanism through KIFC1 inhibition could be exploited in small-cell lung cancer (SCLC), an aggressive malignancy with limited treatment options and poor prognosis. Through in silico and in vitro analyses, as well as IHC of clinical samples, we found that KIFC1 is overexpressed and that centrosome amplification occurs more frequently in SCLC compared with normal tissues and other cancer types. Pharmacological and genetic inhibition of KIFC1 disrupted the clustering of supernumerary centrosomes, triggered multipolar mitosis, and exerted antineoplastic effects in SCLC cells, with minimal effects on noncancerous cells. These findings were validated and extended in vivo using SCLC xenograft models. Finally, cotargeting KIFC1 and the centrosome duplication regulator PLK4 further enhanced growth suppression in SCLC cells. Together, these results suggest that disrupting centrosome clustering and triggering anaphase catastrophe via KIFC1 inhibition may represent a promising therapeutic strategy for SCLC.
Authors
Natsuki Nakagawa, Minemichi Toda, Akiko Kunita, Masafumi Horie, Masakatsu Tokunaga, Hiroaki Ikushima, Mirei Ka, Takahiro Iida, Manabu Shigeoka, Yukinobu Ito, Takahiro Ando, Kousuke Watanabe, Yasunori Ota, Xi Liu, Ethan Dmitrovsky, Hidenori Kage, Masanori Kawakami
No posts were found with this tag.